Bandage
Bioinformatics app for navigating de novo assembly graphs
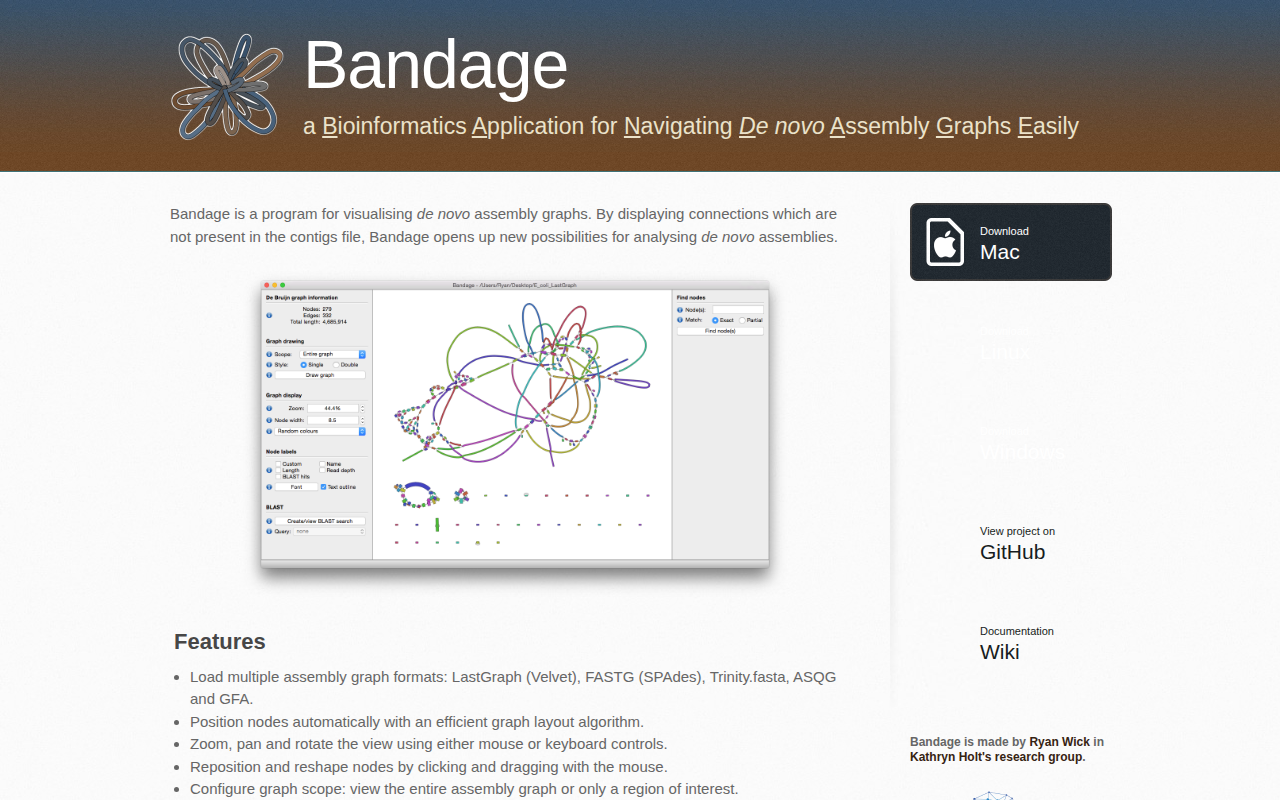
The application visualises de novo assembly graphs, showing connections that are absent from contig files and thereby extending the analyst’s view of assembly structure. It supports several graph formats—including LastGraph, FASTG, Trinity.fasta, ASQG and GFA— and automatically positions nodes with an efficient layout algorithm. Users can navigate the graph by zooming, panning, rotating, or dragging nodes, and can limit the view to the whole assembly or a specific region of interest.
Features include copying or saving node sequences, colour‑coding nodes with built‑in or custom schemes, and labelling by number, length, coverage or user‑defined tags. Nodes can be displayed in single‑ or double‑node styles, with thickness reflecting read depth, and integrated BLAST search highlights and labels specific sequences. Command‑line access allows batch loading of settings, and image generation.
The tool is intended for bioinformaticians who need to assess assembly quality, locate problematic regions, resolve ambiguities, and extract candidate sequences spanning multiple nodes. It runs on macOS, Linux and Windows as a portable 64‑bit binary requiring no installation, with optional source builds for custom environments.
Reviews
Loading reviews…
Similar apps
Databases & Data Tools
Integrative Genomics Viewer (IGV)
Visual exploration of genomic data
Documents, Forms & Contracts
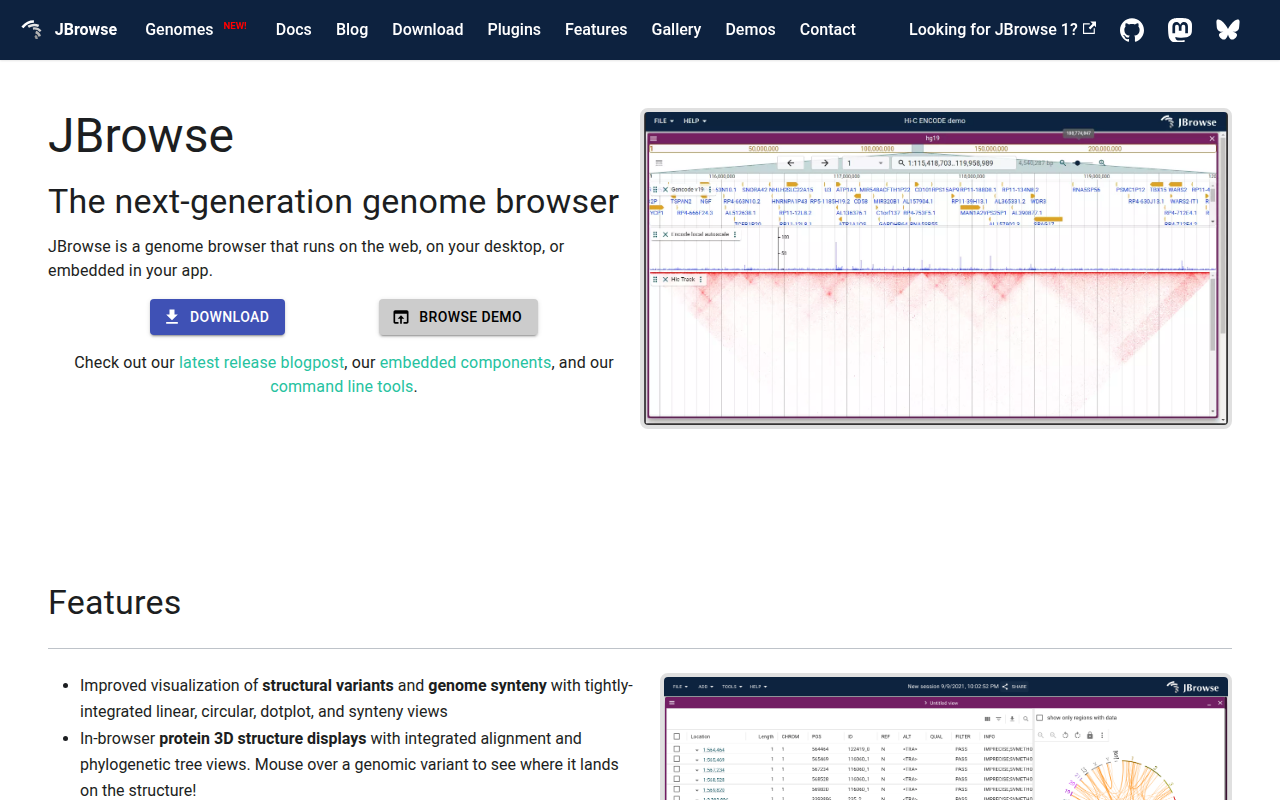
JBrowse
Genome browser
Code Editors & IDEs
ApE (A Plasmid Editor)
Software for DNA sequence analysis and annotation
Databases & Data Tools
Geneious Prime
Bioinformatics software platform
Documents, Forms & Contracts
Jalview
Multiple sequence alignment editor, visualiser, analysis and figure generator
System Monitoring & Maintenance
TreeViewer
Phylogenetic tree viewer
