JBrowse
Genome browser
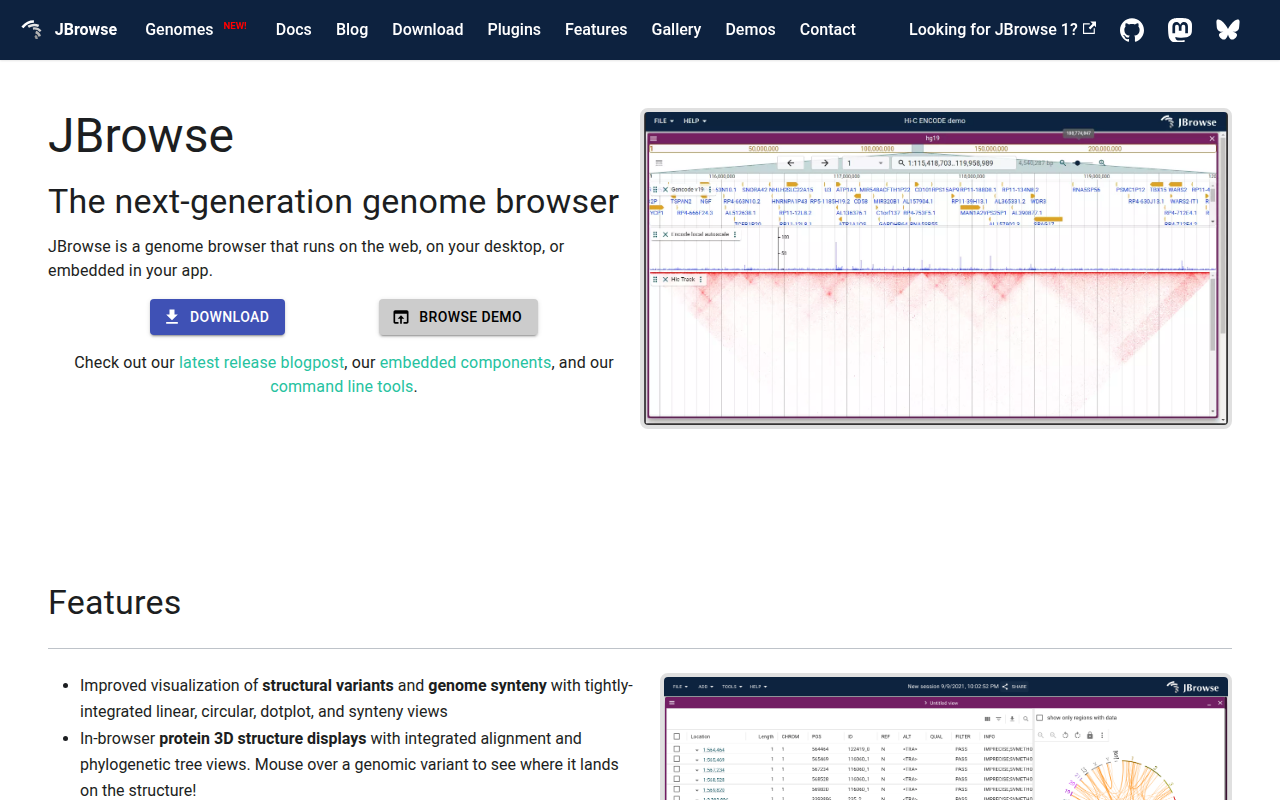
JBrowse is a web‑based genome browser that can also run on a desktop or be embedded in other applications. It visualizes a wide range of genomic data, including structural variants, synteny, and protein 3‑D structures, and offers linked linear, circular, dot‑plot, and phylogenetic tree views. Users can hover over variants to see their position on protein structures, and the tool supports common formats such as BAM, CRAM, VCF, GFF, BED, BigBed, and BigWig.
The platform is built for extensibility; a plugin ecosystem lets developers add new view types, track types, and data adapters, turning JBrowse into a modular computational environment for integrating diverse biological datasets. It is released under the Apache License 2.0 and is maintained by the JBrowse Consortium, which reports research citations as a metric of impact.
JBrowse targets researchers and bioinformaticians who need an interactive, customizable interface for exploring genomic data across multiple formats and visual representations, and it is supported by funding from NIH, the Chan Zuckerberg Initiative, and other research institutions.
Reviews
Loading reviews…
Similar apps
Databases & Data Tools
Integrative Genomics Viewer (IGV)
Visual exploration of genomic data
Documents, Forms & Contracts
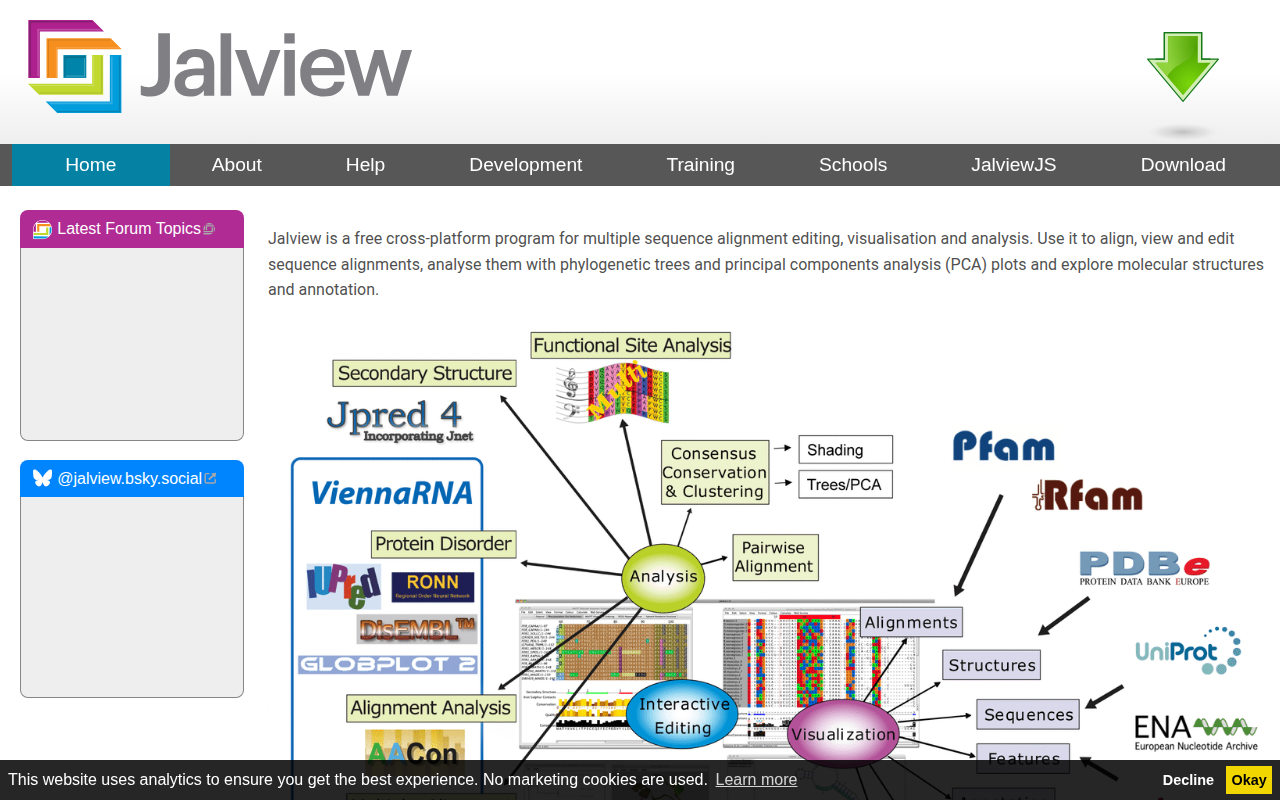
Jalview
Multiple sequence alignment editor, visualiser, analysis and figure generator
Databases & Data Tools
Geneious Prime
Bioinformatics software platform
Documents, Forms & Contracts
jamovi
Statistical software
Code Editors & IDEs
Ugene
Free open-source cross-platform bioinformatics software
Code Editors & IDEs
ApE (A Plasmid Editor)
Software for DNA sequence analysis and annotation
