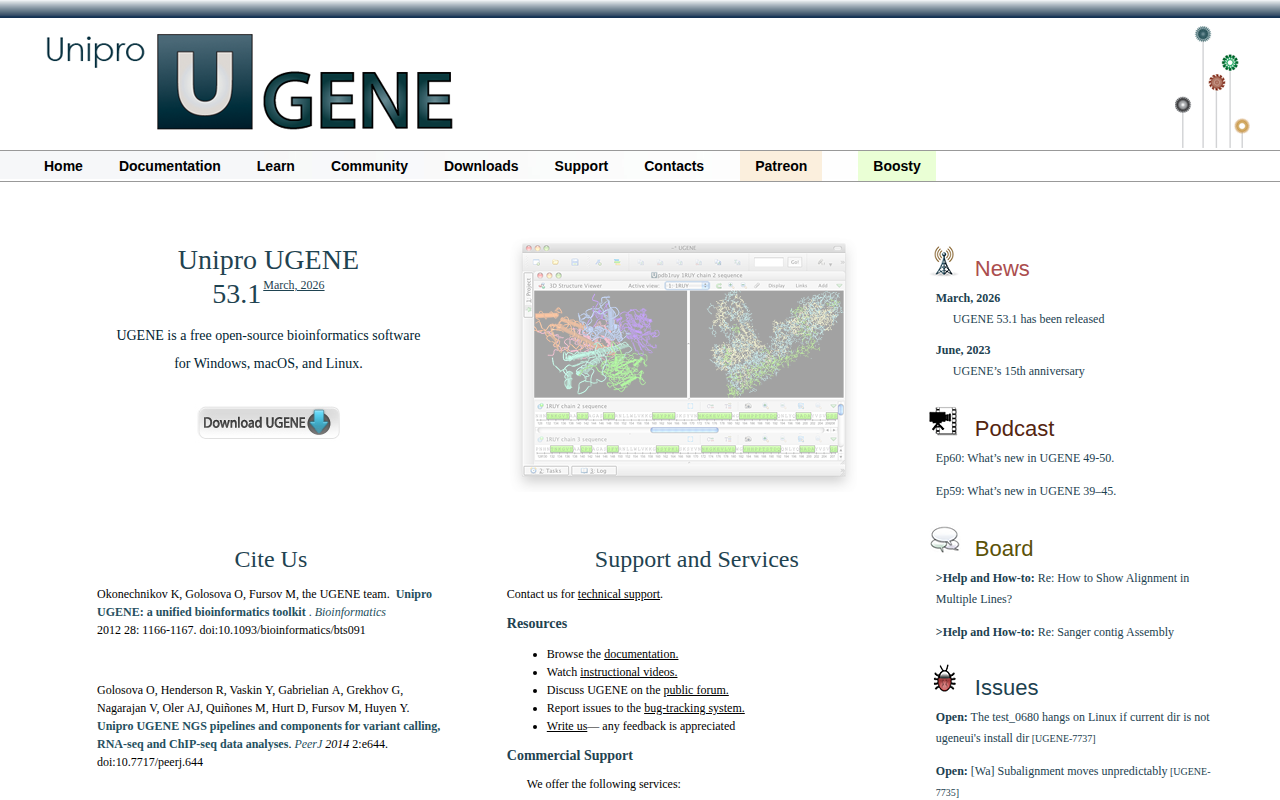
Ugene
Free open-source cross-platform bioinformatics software
The software provides a unified environment for visualizing DNA and protein sequences, performing alignments, assembling genomes, and annotating genetic data. It supports common bioinformatics workflows such as variant calling, RNA‑seq, ChIP‑seq, and metagenomics classification, offering built‑in pipelines and components for these analyses. Users can view alignments in multiple formats, construct custom pipelines, and automate tasks through a graphical interface.
It is aimed at researchers, bioinformaticians, and laboratory technicians who need a cross‑platform tool for sequence analysis without requiring separate specialized applications. The program runs on Windows, macOS, and Linux, and includes documentation, instructional videos, and a public forum for community support. Commercial services are available for custom feature development, defect fixing, and deployment on remote servers.
The project is open source and has been maintained for over a decade, with regular releases and peer‑reviewed publications describing its architecture and applications. Its stable maturity level indicates that core functionality is reliable for routine bioinformatics work.
Reviews
Loading reviews…
Similar apps
Databases & Data Tools
Geneious Prime
Bioinformatics software platform
Databases & Data Tools
Integrative Genomics Viewer (IGV)
Visual exploration of genomic data
Code Editors & IDEs
ApE (A Plasmid Editor)
Software for DNA sequence analysis and annotation
Code Editors & IDEs
MEGA
Molecular evolution statistical analysis and construction of phylogenetic trees
Documents, Forms & Contracts
JBrowse
Genome browser
Documents, Forms & Contracts
Jalview
Multiple sequence alignment editor, visualiser, analysis and figure generator
